Huang a Review of Computational Methods for Predicting Drug Targets
- Enquiry
- Open Admission
- Published:
An efficient computational method for predicting drug-target interactions using weighted extreme learning machine and speed upwards robot features
BioData Mining book xiv, Article number:3 (2021) Cite this article
Abstract
Background
Prediction of novel Drug–Target interactions (DTIs) plays an important part in discovering new drug candidates and finding new proteins to target. In consideration of the time-consuming and expensive of experimental methods. Therefore, information technology is a challenging task that how to develop efficient computational approaches for the authentic predicting potential associations betwixt drug and target.
Results
In the paper, we proposed a novel computational method called WELM-SURF based on drug fingerprints and protein evolutionary information for identifying DTIs. More specifically, for exploiting protein sequence feature, Position Specific Scoring Matrix (PSSM) is applied to capturing protein evolutionary data and Speed upward robot features (SURF) is employed to excerpt sequence key feature from PSSM. For drug fingerprints, the chemical structure of molecular substructure fingerprints was used to represent drug as feature vector. Have business relationship of the advantage that the Weighted Extreme Learning Automobile (WELM) has short training time, proficient generalization power, and almost importantly ability to efficiently execute classification by optimizing the loss part of weight matrix. Therefore, the WELM classifier is used to deport out nomenclature based on extracted features for predicting DTIs. The performance of the WELM-SURF model was evaluated by experimental validations on enzyme, ion channel, GPCRs and nuclear receptor datasets past using fivefold cantankerous-validation test. The WELM-SURF obtained average accuracies of 93.54, ninety.58, 85.43 and 77.45% on enzyme, ion channels, GPCRs and nuclear receptor dataset respectively. We too compared our operation with the Farthermost Learning Motorcar (ELM), the land-of-the-art Support Vector Machine (SVM) on enzyme and ion channels dataset and other exiting methods on four datasets. By comparison with experimental results, the performance of WELM-SURF is significantly better than that of ELM, SVM and other previous methods in the domain.
Conclusion
The results demonstrated that the proposed WELM-SURF model is competent for predicting DTIs with loftier accuracy and robustness. It is predictable that the WELM-SURF method is a useful computational tool to facilitate widely bioinformatics studies related to DTIs prediction.
Background
The noesis of drug-target interactions (DTIs) is much essential for drug evolution, and so more and more studies take pay attending to identify drug-target interactions (DTIs). Identifying of novel DTIs can provide a certain help in drug development and finding new target proteins and discovering new drug candidates [1, 2]. In contempo years, many experimental methods accept been adult for identifying associations between drug and target protein, nevertheless, which are expensive and time-consuming. Developing a successful new chemistry-based drug usually costs billions of dollars, and information technology takes nearly a decade to bring the drug into market. All the same, merely a few drug candidates are approved for marketing past Food and Drug Administration (FDA) [three,4,five]. The major reason is that lack of cognition of DTIs, resulting in unacceptable toxicity for drug candidates. Nonetheless, more and more studies have shown that the DTIs can provide a significant effect on the toxic side effects or toxicity of drug compounds. The knowledge of protein-target interactions can provide a certain aid in finding the toxicity of drug candidates [6]. In addition, identifying interactions between poly peptide and target can also help detecting new potential targets for an old drug and finding many potential drug candidates for a new drug target. Identifying of all potential targets could bring about a better agreement of potential toxicity and handling of other diseases. Because of the inherent disadvantages of experimental methods, it is an urgent chore for developing efficient computational approaches to identify DTIs. As a consequence, using computational approaches for predicting DTIs is becoming more and more than of import. New potential drug–target interaction candidates could be discovered past using computational methods. This make information technology can reduce the price and time of experimental methods and provide a useful validation for biological experimental.
With the completion of the human genome project and the advent of molecular medicine, and with the rapid development of computer applied science and biotechnology, the number of biological science and chemical science biomedical literature is growing rapidly. This enables researchers to restudy the problem related to DTIs through organisation integration. In lodge to computational predict DTIs, many related databases accept been established, some of which are freely available from the public sector and pay attending to relationships between drug and target, for example, Kyoto Encyclopedia of Genes and Genomes (KEGG) [7] SuperTarget and Matador [8], DrugBank [9, ten] and Therapeutic Target Database (TTD) [xi, 12]. The about of import aid is that the information stored in these databases tin provide an amount of essential experimental materials for researchers to develop new computational methods for detecting DTIs on large-scale and widely genome.
Because of the importance of identifying DTIs, a large number of computational approaches have been presented to find DTIs. These methods can exist classified equally ii categories: the ligand-based virtual screening approach and docking simulation. The first method compares the similarity of a given poly peptide based on chemical structure with a classical SAR framework to predict DTIs [thirteen]. However, this method has the disadvantage of not using protein domain information. The second method is a very useful tool of molecular modeling, which can detect the positive interactions between drug molecules and proteins by dynamically simulating the binding between drug molecules and proteins [14,xv,16]. However, the method has a significantly disadvantage that information technology can be merely applied to the proteins of known 3D protein structure. So far, all proteins just comprise a fraction of the proteins of known 3D protein structure, therefore, the Docking simulation method is hard to meet the experimental atmospheric condition. In addition, compared with the data of known 3D protein structure, more and more protein sequence data have been detected, and the poly peptide sequence data are increasing exponentially. Therefore, information technology is very urgent inquiry for develop efficient computational approaches based on poly peptide sequence to identify DTIs.
Recently, a large number of computational methods take been developed to identify DTIs. Yang et al [17] proposed a computational method for finding optimal multi-objective intervention schemes in disease networks. For better recovering the disease network to the desired normal state, the method attempts to identify effective intervention points and combinations of interventions in a given disease network. Kun-Yi Hsin et al [18] proposed a new computational method, which combines two machine learning models carefully developed with multiple docking packages to evaluate the binding potential of a exam compound to proteins involved in complex molecular networks. The prediction model obtained good prediction results. Francisco et al [19] presented a approach for identifying DTIs, which used molecular 2D descriptors to generate drug feature vectors. Chen et al [20] adult an effective classifier to find DTIs by integrating the chemic-poly peptide connections information and chemical-chemical similarities information. Yan et al [21] proposed a new feature extraction method, which used the similarity of drug chemical and target protein sequence to represent drug-target pairs. The random woods was employed to carry out prediction. Zhang at el [22] proposed a ensemble learning algorithm to boost performance of previous DTIs prediction methods through employing the SVM classifier to integrate the prediction results of previous methods. In spite of this, it is very of import for researchers to develop efficient and robustness computational methods for improving prediction accuracy of identifying DTIs.
In the paper, nosotros proposed a novel computational method called WELM-SURF based on drug fingerprints and poly peptide evolutionary information for identifying DTIs. More than specifically, for exploiting protein sequence characteristic, Position Specific Scoring Matrix (PSSM) is applied to capturing protein evolutionary information and Speed up robot features (SURF) is employed to extract sequence fundamental feature from PSSM. For drug fingerprints, the chemical construction of molecular substructure fingerprints was used to represent drug every bit feature vector. Take account of the reward that the Weighted Farthermost Learning Machine (WELM) has short training time, expert generalization ability, and most chiefly ability to efficiently execute classification by optimizing the loss function of weight matrix. Therefore, the WELM classifier is used to deport out nomenclature based on extracted features for predicting DTIs. The performance of the WELM-SURF model was evaluated by experimental validations on enzyme, ion channel, GPCRs and nuclear receptor datasets by using fivefold cross-validation examination. The WELM-SURF obtained average accuracies of 93.54, 90.58, 85.43 and 77.45% on enzyme, ion channels, GPCRs and nuclear receptor dataset respectively. We as well compared our operation with the Extreme Learning Machine (ELM), the state-of-the-art Back up Vector Automobile (SVM) on enzyme and ion channels dataset and other exiting methods on four datasets. By comparing with experimental results, the performance of WELM-SURF is significantly ameliorate than that of ELM, SVM and other previous methods in the domain. The results demonstrated that the proposed WELM-SURF model is competent for predicting DTIs with loftier accurateness and robustness. It is anticipated that the WELM-SURF method is a useful computational tool to facilitate widely bioinformatics studies related to DTIs prediction..
Method
Datasets
In the work, we evaluate the performance of the WELM-SURF model on four datasets: enzymes, ion channels, GPCRs and nuclear receptors. They can be downloaded from the KEGG BRITE [7], BRENDA [23], SuperTarget [8] and DrugBank [9] databases and defined as the golden standard datasets past Yamanishi [24]. The number of known drugs for enzymes, ion channels, GPCRs and nuclear receptors are 445, 210, 233 and 54 and the count of known target protein are 664, 204, 95 and 26. Afterwards advisedly screening, 5127 drug-target pairs tin can interact with each other. In that location are 2926, 1476, 635, and 90 known interactions involving enzymes, ion channels, GPCRs, and nuclear receptors. Therefore, we synthetic positive samples for each of the four datasets.
Usually, a bipartite graph was used to represent Drug-target interaction network, where each node stand for drug molecules or target protein, and each border describes true drug-target interactions valeted by biological experiments or other methods. As can be seen from the bipartite graph, the numbers of existent drug-target interactions edges are small [25]. Hither, nosotros take the enzyme dataset as an example, there are 295,480 connections (445 × 664) in the corresponding bipartite graph, of which only 2926 edges are known drug-target interactions. Therefore, the possible count of negative samples (295480–2926 = 29,255) is significantly larger than the number of positive samples (2926). As a result, this will lead to a bias trouble. For addressing this problem, we randomly selected the aforementioned number of negative and positive samples. Therefore, the number of negative samples for the enzyme, ion channel, GPCRs, and nuclear receptor are 2926, 1476, 635, and 90, respectively. In fact, at that place may be the real drug-target pairs among these negative sample sets. Yet, take business relationship of the large scale of the bipartite graph, the number of true interaction pairs defined as the negative pairs is very small.
Characteristic extraction
Drug molecules description
Recently, a number of biological experiments have indicated that drugs with like chemical structure have similar therapeutic functions. In order to represent drugs as feature vectors, several kinds of descriptors have been designed, such every bit, molecular substructure fingerprints, ramble, topological, geometrical and quantum chemical properties. In the paper, the chemical structure of molecular substructure fingerprints was used to stand for the drugs every bit drug characteristic vectors [15]. Each molecular structure is translated into a fingerprint of a structural by using the molecular fingerprints method. This make information technology can be divers as an 881 dimensional binary vector and its respective bits is 1 or 0.
Position specific scoring matrix (PSSM)
Due to proteins are functionally conserved, the prediction performance can be improved by using the evolutionary data of poly peptide sequence. The position-specific scoring matrix (PSSM) contains not only the position information of the protein sequence, but also the development information that reflects the conservative function of protein. In the experiment, each protein sequence was converted a L × 20 PSSM by using Position Specific Iterated BLAST (PSI-Smash) tool [26], where 50 represents the length of unlike protein sequences. Therefore, nosotros employed the PSSM for extracting the sequence evolutionary information because of its advantage in the paper. The diagram of PSSM is displayed in Fig. 1.
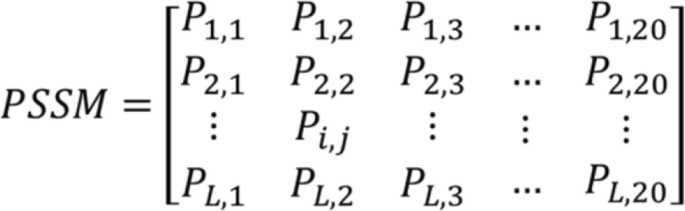
The diagram of PSSM
Where 20 are twenty different amino acids, P ij represent the probability that the i th amino acid in the sequence is mutated to the j th type amino acid during biological evolution. The P ij is greater than 0, equal to 0 and less than 0. If the P ij is a positive number that indicates the i th amino acid can be easily mutated to the jth amino acrid. In practise, the larger number of P ij means a college mutation probability. Conversely, if P ij is negative number, it means the mutation probability is pocket-sized, and a smaller P ij number indicates more conservative. For using evolutionary information of poly peptide sequences to capture more key features, we converted each poly peptide sequence into a PSSM through employing PSI-Smash tool. In the experiment, we set the parameter of PSI_BLAST'south e-value is 0.001 and selected three iterations for obtaining widely and highly homologous sequences.
Speed up robot features (SURF)
Speed up robot features (SURF) [27] characteristic extraction algorithm is an improvement of Scale Invariant Feature Transform (SIFT) algorithm [28, 29], which runs faster than SIFT in algorithm execution efficiency. The SIFT uses Gaussian differences to estimate Laplace Gauss distribution to find calibration space. Withal, the SURF uses Box Filter to approximate LOG. The major advantage of SURF is that it is easier to calculate the convolution with the box filter by using the integrated epitome, which tin be washed in parallel at different scales. The execution of the SURF algorithm depends on the determinant of the Hessian matrix and the determinant of the position. The SURF algorithm includes the following two steps: feature signal detection and feature adjacent description.
Characteristic point detection
The SURF uses continuous Gaussian filters of different scales to process prototype and detects feature points of mesoscale invariant through Gaussian differences. SURF can represent Gaussian fuzzy approximation by using the square filter to supplant the Gaussian filters of SIFT. The SURF feature extraction approach can convert a image into sets of vectors I J ∈R d , j = 1, …, N, where N is a number of images and I j = {f 1,f two, …f m } and \( {f}_m=\left\{{f}_m^1,{f}_m^ii,\dots .{f}_m^d\right\} \) , where 1000 is a number of local features in each image and d is the SURF features dimension that is equal to 64. The fm correspond the SURF local features, note that the m values are not same in all images. We want to organize I j into Grand clustersc = {c 1,c 2, …c k }. The similarity criterion then is defined as follow equation:
$$ S\left(x,y\right)=\sum \limits_{i=1}^chiliad\sum \limits_{j=i}^due north{a}_i^j sim\left({I}_j,{m}_j\right) $$
Where \( 10=\left\{{a}_i^j\right\} \) isouthward separation matrix, \( {\mathrm{a}}_{\mathrm{i}}^{\mathrm{j}}=\left\{\brainstorm{array}{c}i,\kern0.5em \mathrm{if}\ {\mathrm{a}}_{\mathrm{i}}^{\mathrm{j}}\in \mathrm{clusters}\\ {}0,\kern0.5em \mathrm{otherwise}\end{array}\correct. \) with \( {\sum}_{i=1}^k{\sum}_{j=1}^n{a}_i^j=1\forall j;y \) = { m i, …, m k }, sim(I j ,m j ) represents how the correspondent features can exist calculated between the two sets of local features.
The square filter can greatly improve the computation speed through using integral graph that only calculates the value the four corners of the square filter. The determinant value of hessian matrix represents the change around pixel points. Since SURF USES hessian matrix of spot detection to identify characteristic point whose value should exist divers as the maximum or minimum value of determinant. In addition, in guild to achieve scale invariance, SURF likewise USES the determinant of calibration σ to deport out detection of feature signal. For case, given a bespeak p = (x, y) in the graph, the Hessian matrix of scale σ is tin be represented as follows:
$$ H\left(p,\sigma \correct)=\left(\brainstorm{array}{c}{L}_{xx}\left(p,\sigma \right)\kern1.5em {L}_{xy}\left(p,\sigma \right)\\ {}{L}_{xy}\left(p,\sigma \right)\kern1.5em {L}_{yy}\left(p,\sigma \right)\end{assortment}\correct) $$
Where the Lxx(p, σ) , 50xy(p, σ), Fiftyxy(p, σ) and 50yy(p, σ) are the grey-social club epitome later the 2d order differentiation. The Calibration of SURF isn't continuous Gaussian ambiguity and down sampling processing. On the opposite, it is determined past the size of foursquare filters. The lowest scale (initial scale) of square filter of is 9 × nine, which is approximately σ =one.ii Gaussian filter. The size of the upper scale filter will get larger and larger, such equally xv × 15, 21 × 21, 27 × 27…
The transformation formula of its calibration is equally follows:
$$ {\sigma}_{approx}= Currentfiltersize\times \left(\frac{BaseFilterscale}{BaseFilterSize}\right) $$
Feature adjacent clarification
The descriptor of SURF uses the concept of Hal wavelet transform. In order to ensure the rotation invariance of characteristic indicate, each characteristic betoken is assigned a management. The SURF descriptors calculate the Hal wavelet transform of 6σ pixels of direction of X and Y around feature point. A vector can exist obtained past add components of corresponding 10 and Y of the wavelet in each interval. The longest (the largest X and Y components) of all vectors is the direction of the feature point. After the direction of the feature point is selected, the descriptor of feature point can exist created by using the direction of surrounding pixels. For example, the 5 × 5 pixel points were defined as a sub region. As a result, a number of xvi sub regions tin can exist generated past extracting the range of 20*xx pixel points effectually the feature point and the ∑dx and ∑dy of the Hal wavelet transform in the X and Y directions within the sub region can be calculated. Finally, a feature vector with dimensional 64 can be generated.
In the experiment, nosotros used SURF method to create characteristic vectors whose dimensional is 64. Figure 2 shows the menstruation diagram of our method.
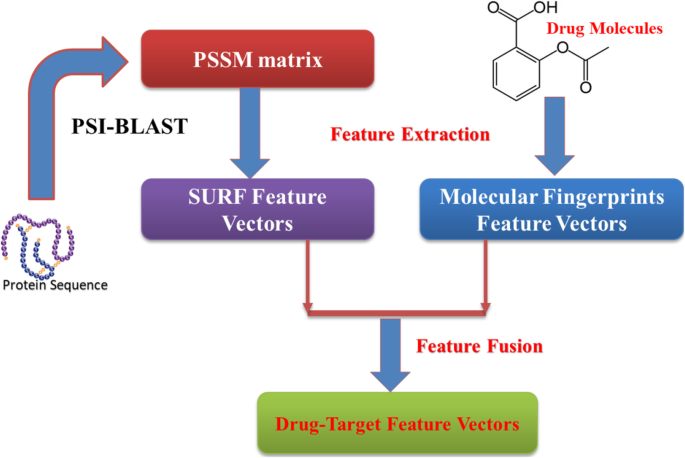
The technology roadmap of the proposed characteristic extraction method
Weighted extreme learning car (WELM)
Zong et al [thirty] proposed a Weighted Extreme Learning Machine (WELM) based on Farthermost Learning Machine (ELM). In gild to efficiently predict DTIs, we build the WELM model based on ELM. The network structure of ELM is equally follows (Fig. 3):
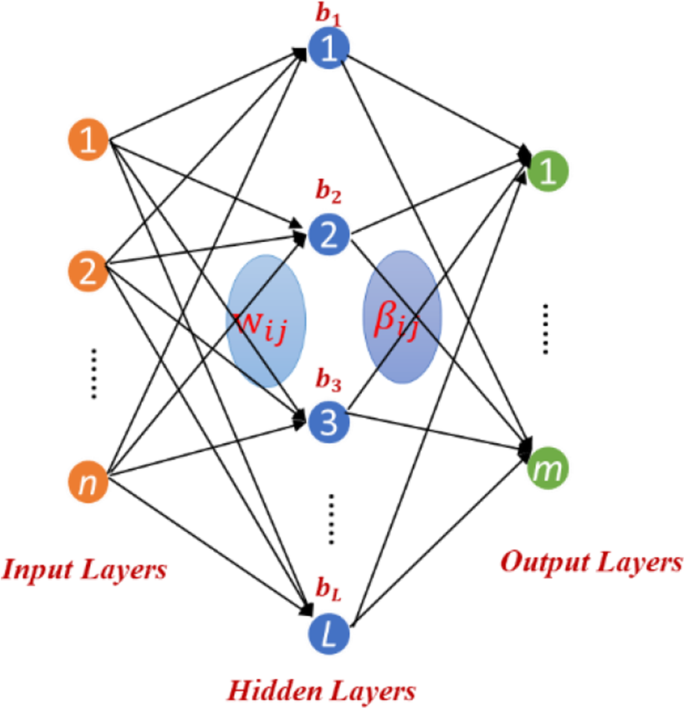
The network structure of ELM
Assuming there are n training samples \( {\left\{{ten}_i,{t}_i\correct\}}_{i=1}^n \), where x i = {x i1,x iii,x i3, ……x in } T ∈R n ,t i = {t i1,t i2,t i3, ……t in } T ∈R thou , n represents the number of sample and 1000 is the classification number. The output model of feedforward neural network with 50 hidden layer nodes can exist expressed as follows:
$$ {\sum}_{h=1}^50{\beta}_hG\left({a}_h,{b}_h,x\right)={o}_i,i=i,2,3,\dots \dots, North $$
(5)
Where β h is the output weight of the h th hidden layer neuron, Yard represents activation role of hidden layer neuron,a h and b h is defined as the input weight and biases of hidden layer neuron,x is input samples, o i represents the bodily output value of i th grooming sample,t i is the expected output of i th training sample. According to the literature [xv], there are N grooming samples \( {\left\{{x}_i,{t}_i\right\}}_{i=one}^north \), x i ∈R n . There are (a h ,b h ) and β h , which make \( {\sum}_{i=1}^L\left|\left|{o}_i-{t}_i\right|\right|=0 \) and unmarried-hidden layer feedforward network (SLFN) can approach the preparation set \( {\left\{{x}_i,{t}_i\correct\}}_{i=one}^north \), ten i ∈R due north with goose egg mistake. The eq. 1 tin can be simplified as follow:
Where H and β are the output matrix and the output weight matrix of the hidden layer respectively and T is the expected output matrix corresponding training samples. The output weight of the subconscious layer can be expressed as follow:
$$ \chapeau{\beta}=\left\{\begin{array}{c}{H}^T{\left(\frac{I}{C}+H{H}^T\right)}^{-1}T,N<L\\ {}{\left(\frac{I}{C}+{H}^Th\right)}^{-i}{H}^TT,N\ge L\end{array}\right\} $$
(7)
The output function of ELM can be defined every bit follow:
$$ f(ten)=h(x)\chapeau{\beta}=\left\{\begin{assortment}{c}h(10){H}^T{\left(\frac{I}{C}+H{H}^T\right)}^{-1}T,N<L\\ {}h(10){\left(\frac{I}{C}+{H}^Th\right)}^{-1}{H}^TT,N\ge Fifty\end{array}\right\} $$
(8)
WELM has two weighting strategies [31], one is automatic weighting and can be defined as follow:
$$ {westward}_1=\frac{one}{Count\left({t}_i\right)} $$
(ix)
Where Count(t i ) represents the number of classt in the preparation sample. The other sacrifices the classification accuracy of the bulk class for obtaining the classification accuracy of the minority class. This splits the minority course and the majority form into 0.618: 1(gold ratio) and is defined as follow:
$$ {w}_2=\left\{\begin{assortment}{c}\frac{0.618}{Count\left({t}_i\right)},{t}_i\in majority\ class\\ {}\frac{1}{Count\left({t}_i\right)},{t}_i\in minority\ form\cease{array}\right\} $$
(x)
The output weight of WELM hidden layer can exist represented as follow:
$$ \hat{\beta}={H}^{-}T\left\{\brainstorm{array}{c}{H}^T{\left(\frac{I}{C}+ WH{H}^T\right)}^{-1} WT,N<50\\ {}{\left(\frac{I}{C}+{H}^T WH\right)}^{-one}{H}^T WT,N\ge L\finish{array}\correct\} $$
(xi)
Where the weighting matrix is a N ×N diagonal matrix, and the N diagonal elements correspond to N samples. Dissimilar weights are assigned to dissimilar sample classes, and the weighting weights of the same class are the same.
The WELM has the advantage of short grooming time and good generalization power and can efficiently execute classification by optimizing the loss role of weight matrix. As a result, the WELM classifier was used to predict DTIs by employing the automatic weighting strategy. The prediction flow diagram of WELM-SURF model is shown in Fig. iv.
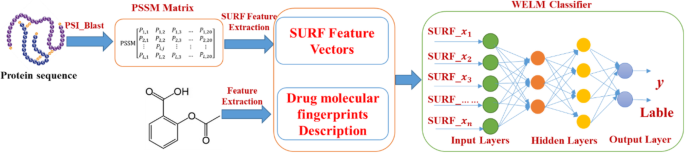
The prediction flowchart of WELM-SURF
Functioning evaluation
The following measures were used to evaleeuate the prediction performance of WELM-SURF in the work.
$$ \mathrm{Acc}=\frac{TP+ TN}{TP+ FP+ TN+ FN} $$
(12)
$$ \mathrm{TPR}=\frac{TP}{TP+ TN} $$
(13)
$$ \mathrm{PPV}=\frac{TP}{FP+ TP} $$
(14)
$$ \mathrm{MCC}=\frac{\left( TP\times TN\right)-\left( FP\times FN\right)}{\sqrt{\left( TP+ FN\right)\times \left( TN+ FP\correct)\times \left( TP+ FP\right)\times \left( TN+ FN\right)}} $$
(15)
Where Acc represents Accuracy, TPR is Sensitivity, PPV is Precision and MCC represents Matthews'south correlation coefficient. TP and TN represent the count of existent interaction and real non-interaction protein sequence pairs correctly predicted. FP and FN is the number of existent non-interaction and real interaction protein sequence pairs mistakenly predicted. Meanwhile, Receiver Operating Curve (ROC) was employed to further assess the prediction performance of WELM-SURF in the work.
Results and discussion
Performance of the proposed method
In the experiment, we evaluate the prediction ability of our WELM-SURF model on four benchmark dataset enzyme, ion channels, GPCRs and nuclear receptor. By and large overfitting will affect experimental results. Therefore, the whole dataset was randomly divided into five parts; four parts were used equally training dataset and the other part was selected as testing dataset. In addition, in society to ensure fairness, fivefold cross-validation tests was employed to evaluate the functioning of the WELM-SURF and several parameters of the WELM model were optimized through using the grid search method. Here, we selected the 'Sigmoid' role and the 'Gaussian 'kernel as the mapping functions of the hidden nodes and gear up Number of Hidden Neurons = 2500, C = 160 and other parameters were prepare up the default value. The prediction results are shown in Tables i, two, 3 and iv using the WELM-SURF prediction model.
It can be observed from Tables 1, two, iii and iv that the average Accurateness for enzymes, ion channels, GPCRs and nuclear receptors is 93.54, 90.48, 85.43 and 77.45% respectively. The corresponding average Sensitivity is 94.58, 91.76, 84.46 and 80.67%, respectively. The corresponding average Precision is 92.86, 89.67, 86.23 and 76.fifty%, respectively. At the same time, the average Matthews's correlation coefficient is 87.89, 82.91, 75.17 and 64.22%, respectively. These experimental results proved that good prediction performance for DTIs prediction tin be achieved past using the WELM-SURF model.
The good experimental results for predicting DTIs are mainly attributed to use the SURF feature extraction method and WELM classifier. The main reward of the WELM-SURF model is that SURF method tin can extract primal evaluation characteristic from PSSM and employed chemical structure of the molecular substructure fingerprints to represent Drug feature and WELM classifier has the advantage of processing sequence data. As discussed, this is mainly due to the following 3 reasons: (i) The PSSM contains not only the position data of the protein sequence, but likewise the development information that reflects the conservative function of protein and a number of prior data. Therefore, information technology can provide a certain help in extracting evolutionary data of protein sequence. Meanwhile, the chemical construction of the molecular substructure fingerprints was use to represent Drug key feature data. (ii) SURF can improve computational speed compared to SIFT. The main advantage of SURF that information technology uses the concept of "calibration infinite" to capture features at multiple calibration levels, which not simply increases the number of available features but as well makes the method highly tolerant to scale changes. This makes it tin can capture DTIs data and extract high efficiency features from PSSM. (3) The WELM has the advantage of short training fourth dimension and practiced generalization power and can efficiently execute classification by optimizing the loss function of weight matrix. Therefore, WELM is used to carry out classification and performs much amend for identifying DTIs in the study. More specifically, the WELM tin can better perceive the distribution information of class by assigning larger weight to the minority class samples and push the separating boundary from the minority course towards the bulk class through using weight strategy. This makes information technology tin can provide help in sensitive learning by assigning different weight. The results demonstrated that the proposed WELM-SURF model tin can better prediction accurateness and is fit for predicting DTIs.
Comparison with the ELM-based and SVM-based method
Despite the proposed WELM-SURF approach obtained adept prediction results. Notwithstanding, in guild to further evaluate the prediction chapters of WELM classifier, we compared its prediction ability with the ELM and the SVM past using SURF characteristic extraction method on enzyme and ion channel datasets. The LIBSVM tool [32] of the SVM was employed to carry out nomenclature. At the same time, for off-white comparison, several parameter of ELM were optimized through employing the aforementioned grid search method. More specifically, the number of subconscious layers of ELM is prepare to 89 and other parameters take the default value. At the same time, the RBF kernel parameters of the SVM were optimized by using the same strategy, where c = 0.half dozen and g = 3.1 and other parameters were gear up the default value.
Table 5, six, 7 and viii list the statistical prediction results of fivefold cantankerous-validation tests on enzyme and ion channels by using ELM classifier and SVM classifier, respectively. At the same time, the comparison of ROC Curves betwixt WELM, ELM and SVM was as well displayed in Fig. v and Fig. 6 on enzyme and ion channels datasets, respectively. It tin be observed from Tables 5 and half-dozen that average accuracy of xc.38 and 87.07% obtained using ELM classifier and SVM classifier on enzyme dataset, while the WELM classifier achieved 93.54% boilerplate accuracy. Similarly every bit shown in Tables 7 and 8, 87.76% average accuracy and 83.thirty% average accuracy are obtained through using ELM classifier and SVM classifier on ion channels dataset. The WELM classifier achieved ninety.48% average accurateness. It tin be seen from comparison results that the prediction capacity of the WELM classifier is significantly better than that of the ELM and the SVM classifier. Similarly, we too can detect from Fig. 5 and Fig. six that the ROC curves of the WELM classifier is too obviously improve than the ELM and the SVM classifier. These good comparison results obtained may be lie in as follows reasons: The significantly advantage of WELM classifier related to the ELM classifier and the SVM Classifier is that it has the advantage of short training fourth dimension and good generalization ability and tin can efficiently execute classification by optimizing the loss function of weight matrix, and can provide a certain help in sensitive learning by assigning different weight. Therefore, experimental results indicated that the proposed prediction model might become useful tools and can identify DTIs with a high prediction accuracy.
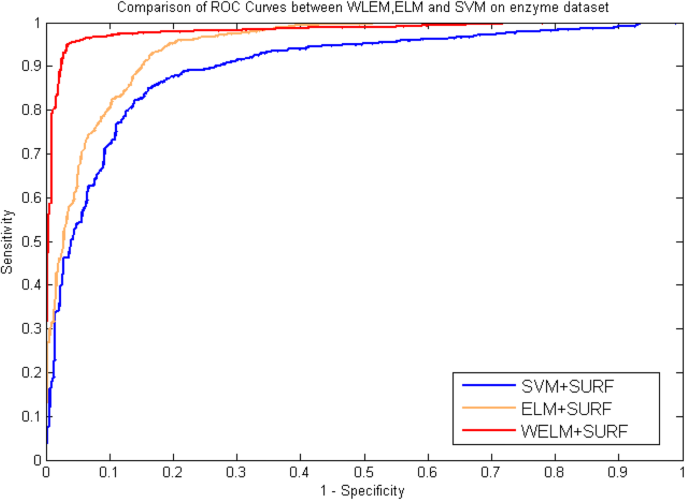
Comparison of ROC curves performed betwixt WELM, ELM and SVM on enzyme

Comparing of ROC curves performed between WELM, ELM and SVM on ion channels
Comparison with other methods
In the paper, for farther evaluating the prediction chapters of WELM-SURF method, we compare our performance with four existing DIIs predictor DBSI [33], Yamanishi [24], KBMF2K [34] and NetCMP [35] on enzyme, ion channels, GPCRs and nuclear receptor dataset. These comparison results are displayed in Tabular array 9. Information technology can be seen from Table 9 that our prediction accurateness is apparently better than that of other four methods. The comparing results are potent show that the WELM-SURF is efficiently and robustness related to current exiting approaches. The results besides demonstrated that the proposed WELM-SURF model is competent for predicting DTIs with loftier accuracy and robustness. It is anticipated that the WELM-SURF method is a useful computational tool and is suitable for predicting DTIs. The primary reason is that the WELM-SURF used a good classifier and developed a novel feature extraction method.
Conclusion
In the paper, we proposed a novel computational method called WELM-SURF, which combines the Weighted Extreme Learning Auto (WELM) with Speeded upward robust features (SURF) to predict DTIs based on drug fingerprints and protein evolutionary information. The WELM-SURF obtained average accuracies of 93.54, 90.58, 85.43 and 77.45% on enzyme, ion channels, GPCRs and nuclear receptor dataset respectively. We also compared our functioning with the ELM classifier and the SVM classifier on enzyme and ion channels dataset and other exiting methods on four datasets. By comparison with experimental results, the operation of WELM-SURF is significantly improve than that of the ELM, the SVM and other previous methods in the domain. This is mainly due to the following three reasons: (ane) The PSSM contains not only the position information of the protein sequence, but too the development information that reflects the bourgeois function of protein and a number of prior information. Therefore, it tin provide a sure assistance in extracting evolutionary information of poly peptide sequence. Meanwhile, the chemical construction of the molecular substructure fingerprints was utilise to stand for Drug key feature information. (two) SURF can improve computational speed compared to SIFT. The main advantage of SURF that it uses the concept of "scale infinite" to capture features at multiple scale levels, which non only increases the number of bachelor features but also makes the method highly tolerant to scale changes. This makes it can capture self-protein interaction information and extract loftier efficiency features from PSSM. (3) The WELM has the advantage of short training fourth dimension and skilful generalization ability and can efficiently execute classification by optimizing the loss office of weight matrix. Therefore, WELM is used to behave out classification and performs much better for identifying DTIs in the study. More specifically, the WELM tin better perceive the distribution information of class by assigning larger weight to the minority form samples and button the separating boundary from the minority class towards the majority class through using weight strategy. This makes it can provide a certain help in sensitive learning by assigning different weight. We can come to the decision that the proposed WELM-SURF model can obtain high prediction accurateness and execute incredibly well for predicting DTIs. For the futurity study, more constructive feature extraction approaches and machine learning algorithms can be adult for predicting DTIs.
Availability of information and materials
In this written report, our experimental datasets can be obtained from the KEGG BRITE [7],BRENDA [23],SuperTarget [8] and DrugBank [9] databases and defined equally the gold standard datasets by Yamanishi [24].
Abbreviations
- DTIs:
-
Drug-target interactions
- WELM:
-
Weighted Extreme Learning Machine
- SIFT:
-
Scale Invariant Feature Transform
- SURF:
-
Speeded Up Robust Features
- PSSM:
-
Position Specific Scoring Matrix
- ELM:
-
Extreme Learning Machine
- SVM:
-
Support Vector Automobile
- PSI-BLAST:
-
Position-Specific Iterated BLAST
- Acc:
-
Accuracy
- TNR:
-
True Negative Rate
- TPR:
-
True Positive Rate
- MCC:
-
Matthews Correlation Coefficient
- PPV:
-
Positive Predictive Value
- ROC:
-
Receiver Operating Curve
References
-
Wang YC, et al. Computationally probing drug-poly peptide interactions via support vector machine. Lett Drug Design Discovery. 2010;seven(5):370–8.
-
Xia Z, et al. Semi-supervised drug-poly peptide interaction prediction from heterogeneous biological spaces. Bmc Syst Biol. 2010;4 Suppl 2(Suppl 2):6.
-
Landry Y, Gies JP. Drugs and their molecular targets: an updated overview. Fundamental Clin Pharmacol. 2008;22(1):ane–18.
-
Li Q, Lai L. Prediction of potential drug targets based on simple sequence properties. Bmc Bioinformatics. 2007;8(1):1–11.
-
Overington JP, Allazikani B, Hopkins AL. How many drug targets are there? Nat Rev Drug Discovery. 2006;5:993.
-
An JY, et al. Computational methods using weighed-extreme learning machine to predict protein self-interactions with protein evolutionary data. J Cheminformatics. 2017;9(1):47.
-
Kanehisa M, et al. From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Res. 2006;34(Database upshot):354–vii.
-
Günther Southward, et al. SuperTarget and matador: resources for exploring drug-target relationships. Nucleic Acids Res. 2008;36(Database consequence):919–22.
-
Wishart DS, et al. DrugBank: a knowledgebase for drugs, drug actions and drug targets. Nucleic Acids Res. 2008;36(Database issue):D901–six.
-
Wishart DS, et al. DrugBank: a comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Res. 2006;34(Suppl 1):668–72.
-
Chen X, Ji ZL, Chen YZ. TTD: therapeutic target database. Nucleic Acids Res. 2002;thirty(1):412–v.
-
Zhu F, et al. Update of TTD: therapeutic target database. Nucleic Acids Res. 2010;38(Database issue):787–91.
-
Butina D, Segall Md, Frankcombe Grand. Predicting ADME properties in silico : methods and models. Drug Discov Today. 2002;seven(11):S83–viii.
-
Coleman RG, Salzberg AC, Cheng Air conditioning. Structure-based identification of small molecule binding sites using a energy model. J Chem Inf Model. 2006;46(vi):2631–7.
-
Cheng AC, et al. Structure-based maximal analogousness model predicts small-molecule druggability. Nat Biotechnol. 2007;25(one):71–5.
-
Sousa SF, Fernandes PA, Ramos MJ. Protein–ligand docking: current condition and future challenges. Proteins Construction Function Bioinformatics. 2006;65(1):xv–26.
-
Yang K, et al. Finding multiple target optimal intervention in disease-related molecular network. Mol Syst Biol. 2008;4(1):228.
-
Hsin, K.Y., et al. Awarding of machine leaning approaches in drug target identification and network pharmacology. in International Briefing on Intelligent Computer science and Biomedical Sciences. 2015.
-
Prado-Prado F, et al. 2D MI-DRAGON: a new predictor for protein–ligands interactions and theoretic-experimental studies of US FDA drug-target network, oxoisoaporphine inhibitors for MAO-A and man parasite proteins. Eur J Med Chem. 2011;46(46):5838–51.
-
Chen X, Yan GY. NRWRH for Drug Target Prediction. in The International Conference on Computational Systems Biological science; 2010.
-
Yan QN. Supervised prediction of drug-target interactions by ensemble learning. J Chem Pharmaceut Res. 2014;6:1991.
-
Zhang R. An Ensemble Learning Arroyo for Improving Drug–Target Interactions Prediction. Cham: Springer International Publishing; 2015. p. 433–42.
-
Schomburg I, et al. BRENDA, the enzyme database: updates and major new developments. Nucleic Acids Res. 2004;32(1):D431–iii.
-
Yamanishi Y, et al. Prediction of drug-target interaction networks from the integration of chemical and genomic spaces. Bioinformatics. 2008;24(xiii):i232–i240(9).
-
Li X, et al. Modulation of gene expression regulated past the transcription factor NF-κB/RelA. J Biol Chem. 2014;289(17):11927–44.
-
Gribskov M, Mclachlan AD, Eisenberg D. Contour analysis: detection of distantly related proteins. Proc Natl Acad Sci U Southward A. 1987;84(13):4355.
-
Bay H, Tuytelaars T, Gool LV. SURF: Speeded Upwardly Robust Features; 2006.
-
Lowe DG. Object Recognition from Local Scale-Invariant Features. in Computer Vision, 1999. The Proceedings of the 7th IEEE International Briefing on; 1999.
-
Lowe DG. Distinctive prototype features from scale-invariant Keypoints. Int J Comput Vis. 2004;60(2):91–110.
-
Zong WW, Huang GB, Chen YQ. Weighted extreme learning machine for imbalance learning. Neurocomputing. 2013;101:229–42.
-
Pan WT. A new Fruit Fly Optimization Algorithm: Taking the financial distress model as an example[J]. Knowledge-Based Systems. 2012;26(ii):69–74.
-
Chang CC, Lin CJ. LIBSVM: a library for support vector machines. Acm Trans Intelligent Syst Technol. 2011;2(3):389–96.
-
Cheng F, et al. Prediction of drug-target interactions and drug repositioning via network-based inference. PLoS Comput Biol. 2012;eight(5):357–72.
-
Gönen M. Predicting drug-target interactions from chemic and genomic kernels using Bayesian matrix factorization. Bioinformatics. 2012;28(18):2304–10.
-
Chen H, Zhang Z. A semi-supervised method for drug-target interaction prediction with consistency in networks. PLoS Ane. 2013;8(5):8750.
Acknowledgments
The authors would like to give thanks all the invitee editors and bearding reviewers for their constructive advices.
Funding
This piece of work is supported by 'the Key Enquiry Funds for the Central Universities (2019XKQYMS88)'.
Author data
Affiliations
Contributions
AJY and MFR conceived the algorithm, carried out analyses, prepared the data sets, carried out experiments, and wrote the manuscript; YZJ and ZYJ designed, performed and analyzed experiments and wrote the manuscript; all authors read and approved the last manuscript.
Respective author
Ethics declarations
Ethics approval and consent to participate
Non applicable.
Consent for publication
Non applicable.
Competing interests
The authors declare no conflict of interest.
Additional information
Publisher'due south Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits utilize, sharing, adaptation, distribution and reproduction in any medium or format, equally long as you lot requite appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were fabricated. The images or other third party cloth in this article are included in the commodity's Creative Eatables licence, unless indicated otherwise in a credit line to the material. If material is not included in the commodity's Creative Eatables licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, yous will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Artistic Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zilch/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the information.
Reprints and Permissions
About this article
Cite this article
An, JY., Meng, FR. & Yan, ZJ. An efficient computational method for predicting drug-target interactions using weighted extreme learning automobile and speed up robot features. BioData Mining fourteen, iii (2021). https://doi.org/x.1186/s13040-021-00242-1
-
Received:
-
Accepted:
-
Published:
-
DOI : https://doi.org/10.1186/s13040-021-00242-1
Keywords
- DTIs
- WELM
- SURF
- PSSM
kearneyhibive1970.blogspot.com
Source: https://biodatamining.biomedcentral.com/articles/10.1186/s13040-021-00242-1
0 Response to "Huang a Review of Computational Methods for Predicting Drug Targets"
Post a Comment